EDITOR'S PICK
-
‘The CCP is the greatest enemy of the Chinese nation’: Statements From the Tuidang Movement (March 2024)
-
Master Li Hongzhi: How Humankind Came to Be
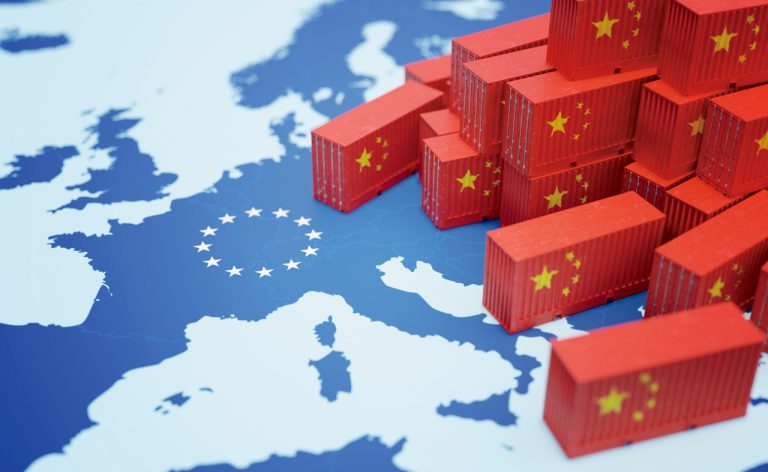

China’s Overseas Product Dumping: Addressing Unfair Competition Caused by Systemic Differences
Faced with unfair competition brought about by systemic differences, countries around the world are right to impose tariffs and other barriers on Communist China
Latest
- For Powerful Traditional Remedies, Know and Grow Medicinal Herbs (G) Geranium
- Caribbean Treasure: Discovering the Rich Heritage of Ron Barceló Imperial Rum
- Mandarin Pop Song Lyrics Shift From Romance to Calls for Freedom, New Research Suggests
- Charges Laid One Year After Historic Canadian Gold Heist
- Hundreds of Migrants Descend on New York City Hall Pleading for Work Permits, Green Cards
FEATURED
-
US Puts Pressure on China as Ukraine War Escalates
-
Communist China Heads Down a Road of Isolation and Impoverishment
-
Chinese-American Artists Targeted in Planned NYT Piece That Would Misrepresent Falun Gong, Shen Yun Performing Arts
-
Shen Yun Artists Face Discrimination from Pro-CCP Official at US Customs Upon Return From European Tour
-
Tibetans in Exile – Raising Voices for a Distant Homeland
-
Toronto Marks 65th Tibetan Uprising Anniversary With Rally Against the CCP
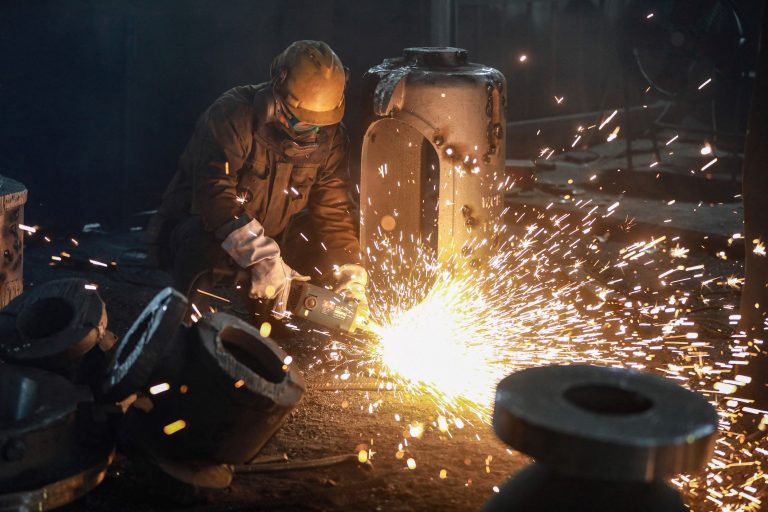

Biden Proposes Tripling Tariffs on Chinese Steel, Aluminum to Counter Beijing’s Product Dumping
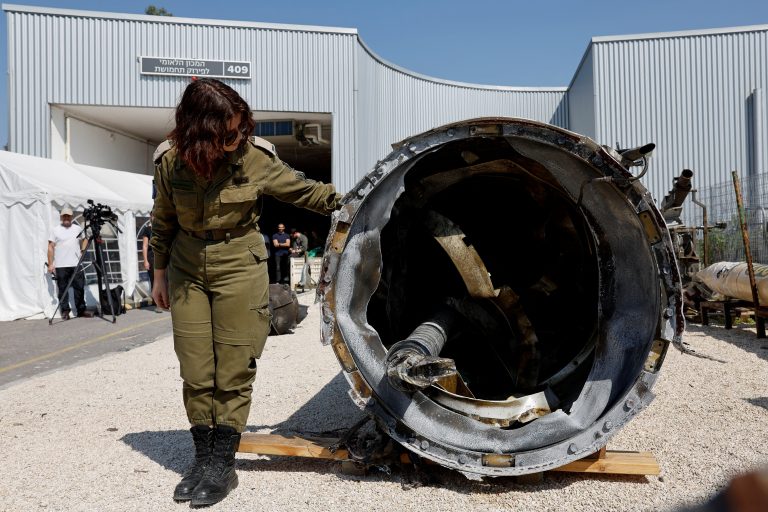

On April 16, U.S. Treasury Secretary Janet Yellen said Iran's attack on Israel last weekend and its financing of militant groups in Gaza, Lebanon, Yemen and Iraq threatened stability in the Middle East and could result in economic spillovers. At a news conference, reading from prepared remarks, Yellen called Iran’s
-
‘Extraordinary artistry’: Shen Yun Delights Theatergoers in Sold-Out Lincoln Center Shows
NEW YORK, New York — After kicking off a highly anticipated 14-show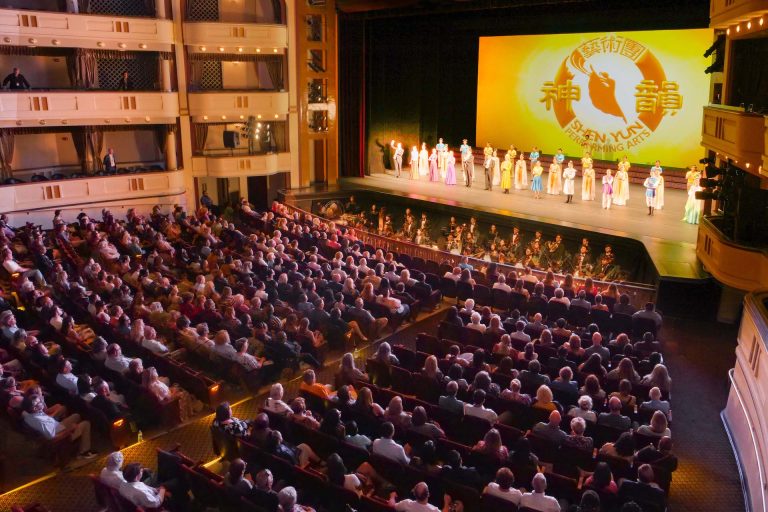
-
‘Very Positive Trends’: Number of Homicides Drops Significantly in the Big Apple
According to numbers compiled by AH Datalytics, the number of nonnegligent manslaughters,
-
Controversial Geoengineering Experiment Launched Quietly in San Francisco to Avoid Public Backlash
On Tuesday, April 2, researchers with the University of Washington quietly conducted
- NY: Long Island Sen. Mario Mattera Tables Bill Addressing Squatting ‘Epidemic’
- Culinary Brilliance on Display at 87 Sussex: Jersey City’s New Dining Gem
- ‘An extension of their souls’: Shen Yun Continues to Wow at Lincoln Center
- ‘Beautiful!’: Shen Yun Wows Audiences in Multiple Sold Out Lincoln Center Performances
- NY: MTA Demands NYC Marathon Pay 750k Congestion Pricing Toll for Using Verrazzano Bridge
- Production Delays, Disarray in Boeing Factory Prior to Door Plug Near Tragedy, WSJ Reports
- White House Directs NASA to Create Time Standard for the Moon
- Trillions of Cicadas to Swarm Several Eastern States This April
- US Officials Celebrate Shen Yun’s Return to Lincoln Center With Letters of Proclamation
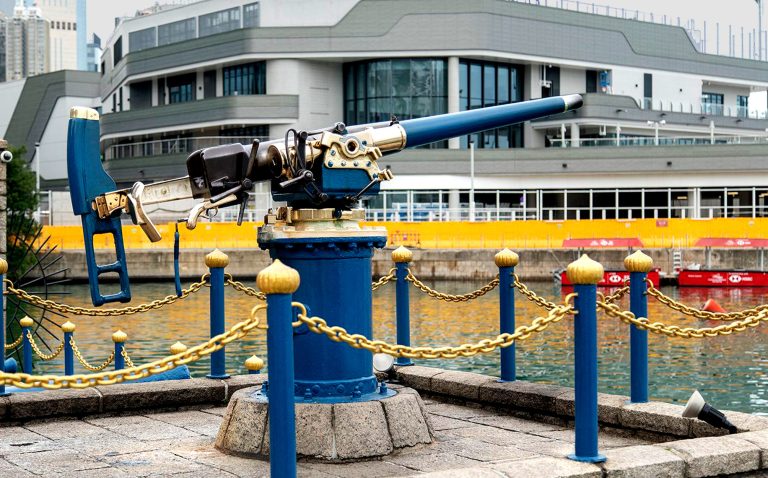

Published with permission from LuxuryWeb Magazine Every day at noon in Hong Kong, a distinct ceremony unfolds — a ship's bell rings followed by the firing of a cannon, its sound reverberating across Causeway Bay. The Noonday Gun has become a cherished ritual, deeply embedded in the city's rich history.
-
How Noncompete Agreements and Corporate Surveillance Are Crushing China’s Tech Workers
In the cutthroat world of China’s tech industry, non-compete clauses and corporate surveillance are increasingly binding tech workers
-
5 Exemplary Women of Ancient China
In ancient China, women were educated and raised to uphold a range of traditional values, which could be
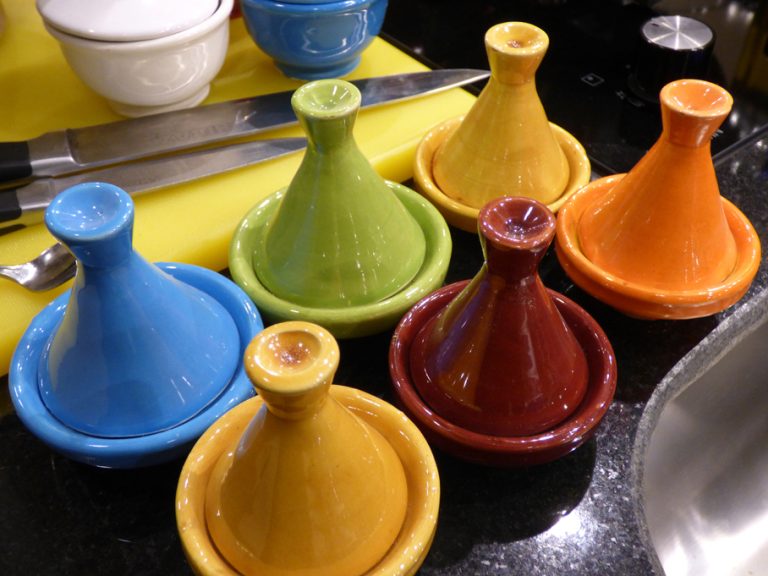

Published with permission from LuxuryWeb Magazine Moroccan culinary arts are steeped in a rich history that spans thousands of years, influenced by the seasonal availability of ingredients, fuel, and cooking implements, as well as a cook’s ambition to create dishes that are delicious, visually appealing, and nutritionally balanced. For instance,
-
Civil War in Sudan Enters 2nd Year: 15,000 Dead, 8 Million Displaced
The civil war in Sudan has been raging for over a year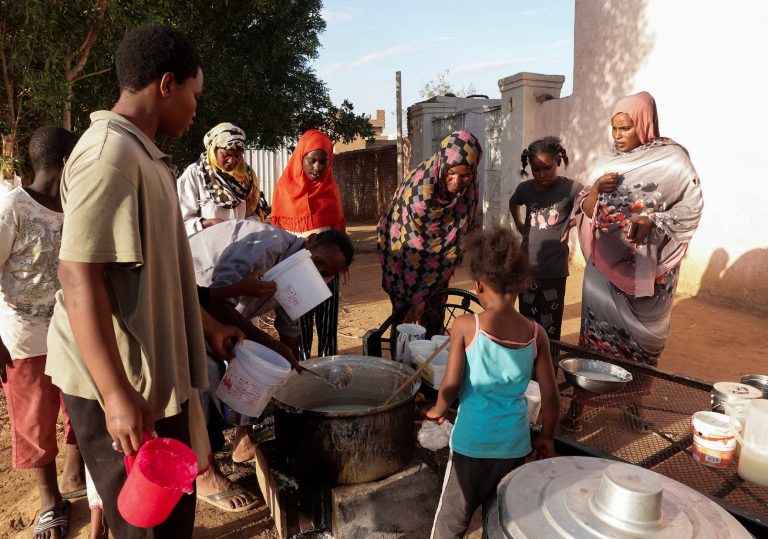
-
Peter Pellegrini Wins Slovak Presidential Election, Beating Pro-Western Opposition
Slovak nationalist government candidate Peter Pellegrini won the country's presidential election on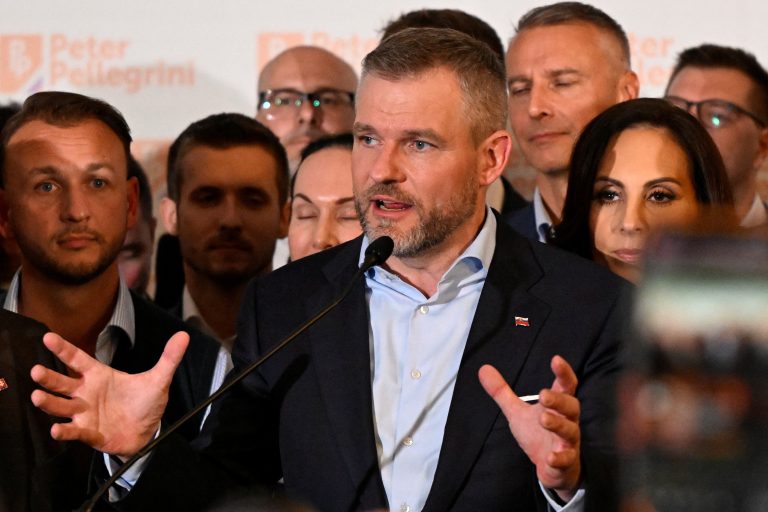
-
Chinese Students Recount Horrifying Experience During Concert Hall Shooting in Moscow
On March 23, several gunmen attacked the Crocus City Hall near Moscow;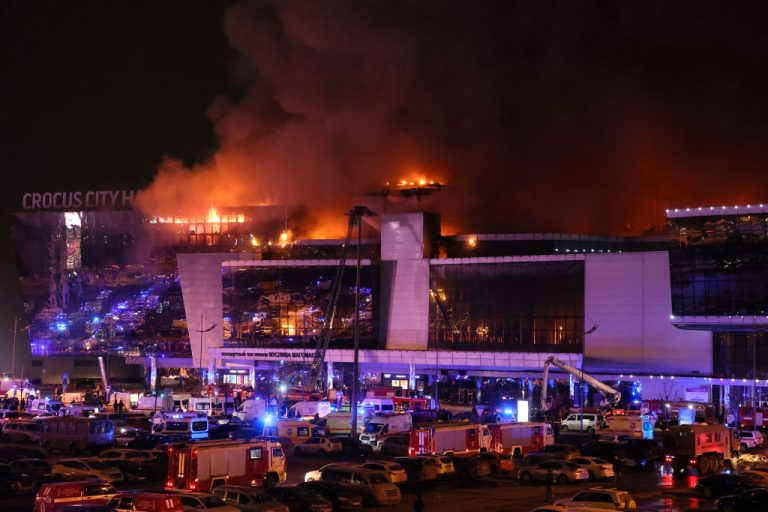
- Japan Reverses Course, Increases Interest Rates for First Time in 17 Years
- Davide Scabin Rumored to Bring Culinary Genius Back to Turin, Italy
- Hong Kong Passes Infamous ‘Article 23’ in Show of Growing Communist Hold
- A Global Guide to Sweet Wines: From Europe to North America
- Trafficked Cambodian Teen Saved Following Facebook Plea
- Argentinian Austerity Measures Show Mixed Results as Inflation Remains High
- US Seeks to Revive Dormant Shipyards With Help From South Korea and Japan
- Sherry: A Divine Nectar From Spain’s Sunny Shores
- Japan’s Daiwa House Planning US Factory for Prefab Homes

In response to a rare 4.8 magnitude earthquake, followed by multiple, smaller aftershocks, many East Coast natives are grappling with earthquake awareness for the first time. Epicentered near Gladstone, New Jersey, slight tremors were felt as far south as northern Virginia and north to the border of New Hampshire. No
-
‘A new breath of life’: Shen Yun’s Opening Night at Lincoln Center Met With Resounding Acclaim
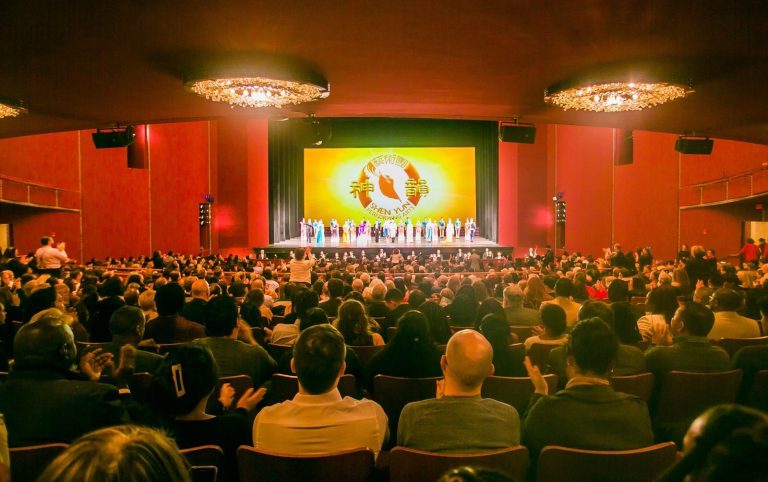 NEW YORK, New York — On April 3, Shen Yun kicked off its highly anticipated 14-show run at the city’s prestigious Lincoln Center. Nearly all tickers were sold out, and theatergoers lucky enough to secure seats raved about the show's vibrant dancing, choreography, gravity-defying acrobatics, and immersive digital backdrops, among
NEW YORK, New York — On April 3, Shen Yun kicked off its highly anticipated 14-show run at the city’s prestigious Lincoln Center. Nearly all tickers were sold out, and theatergoers lucky enough to secure seats raved about the show's vibrant dancing, choreography, gravity-defying acrobatics, and immersive digital backdrops, among
-
A Holistic Approach to Histamine Intolerance
Do you suffer chronic symptoms that can’t be pinned down to any specific cause? Gastrointestinal disorders, headaches, hives,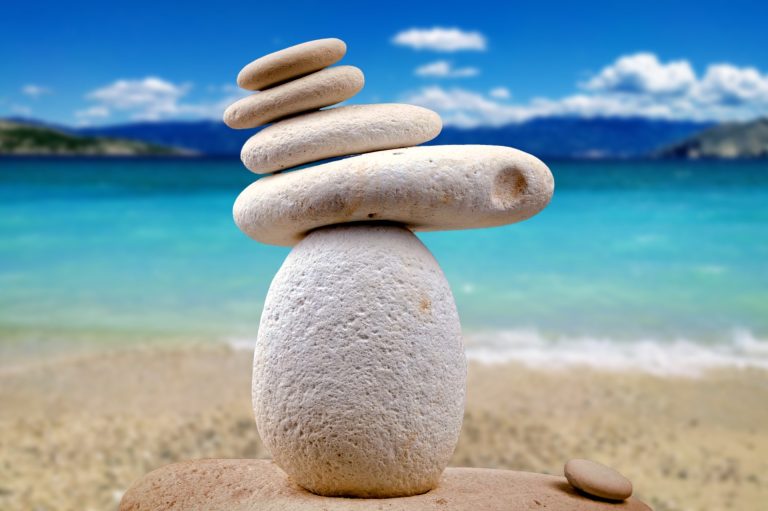
-
‘Absolutely brilliant’: Theatergoers Marvel at Shen Yun Performance in New Jersey
NEW BRUNSWICK, New Jersey — On March 31, Shen Yun concluded a five-show run at the prestigious State
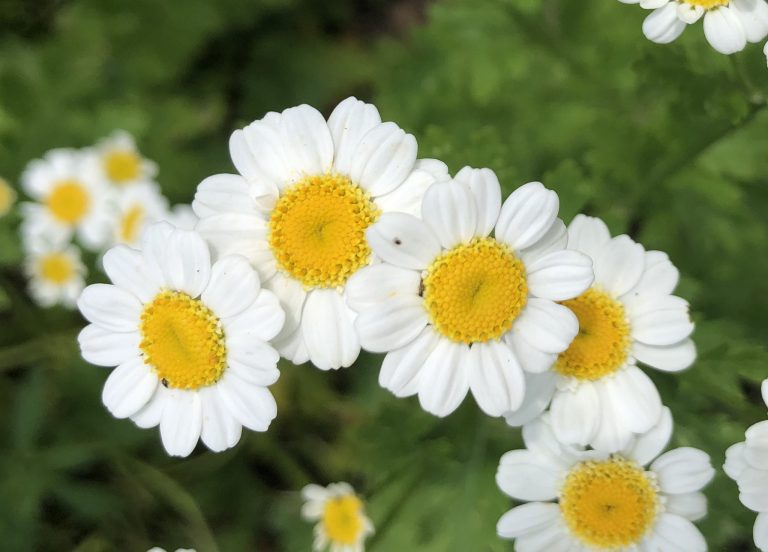

Sprightly and tenacious, feverfew is a member of the world’s largest and most diverse plant family — Asteraceae. Native to Asia and Europe, feverfew was first introduced to the United States in the 19th century. It is now commonly grown as a perennial in hardiness zones 5-10. Its small, prolific
-
The Power of Optimism – The Key to Longevity and Health
Life is hard, and staying positive is even harder. There are those who, despite sporting the scars of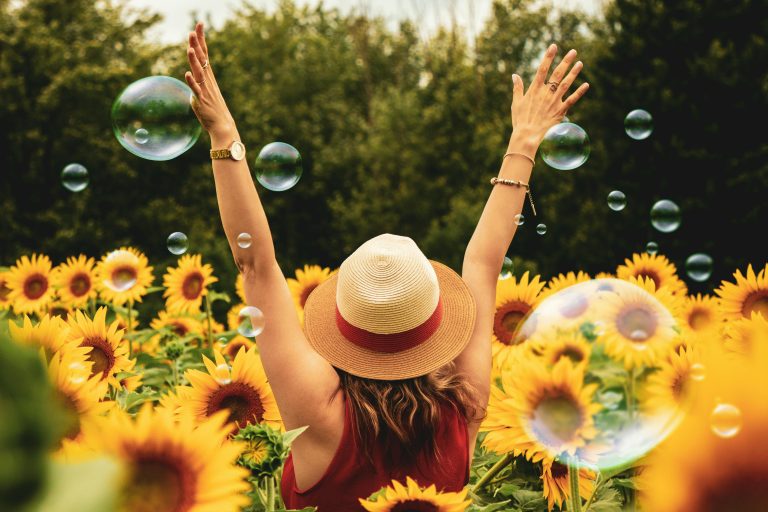
-
Food Allergies – Why We Get Them, and How to Alleviate Them Naturally
Food allergies are an increasingly common dietary fret that can make dining out a risky affair. They occur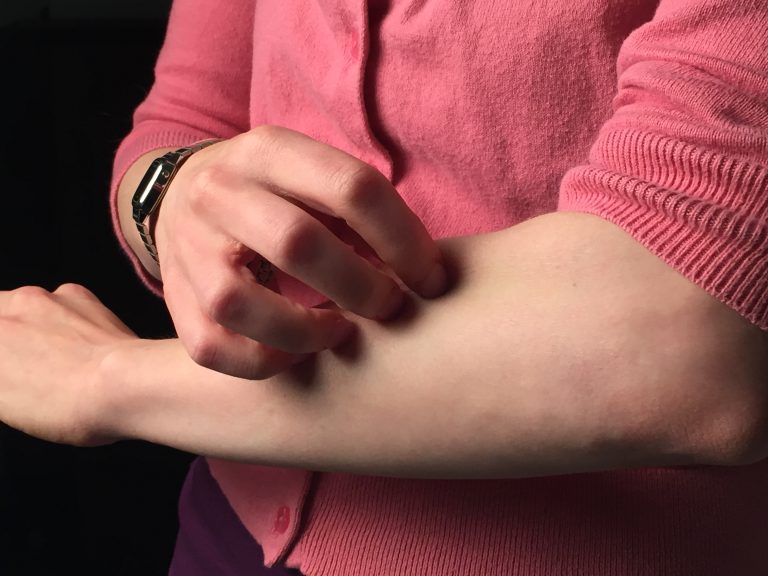

Colorful, artistic and sometimes sweet, decorated Easter eggs are almost a worldwide tradition. These charming gems make their appearance at the beginning of each spring, not only marking a time of rebirth in nature, but also reminding the faithful of Jesus' rise from the tomb.Traditionally, decorated Easter eggs have been
-
Communist China Ramps Up Efforts to Intimidate Shen Yun With Sabotage, Unhinged Threats of Violence
In the most recent manifestation of efforts to sabotage the growing cultural force that is Shen Yun, the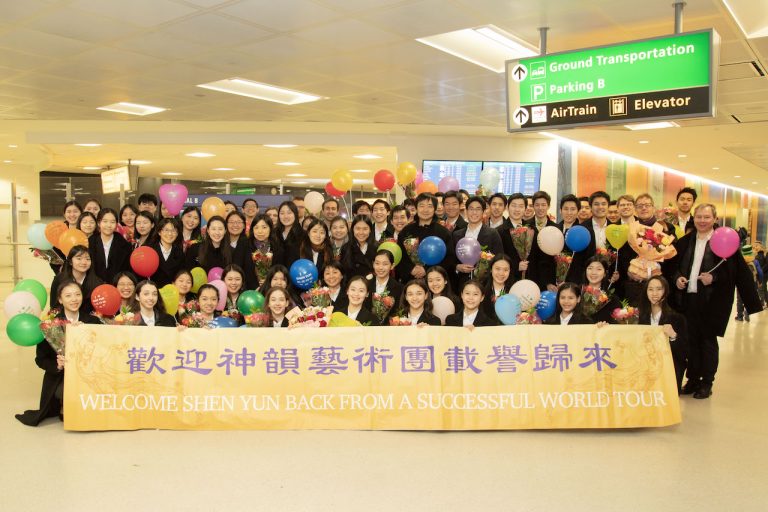
-
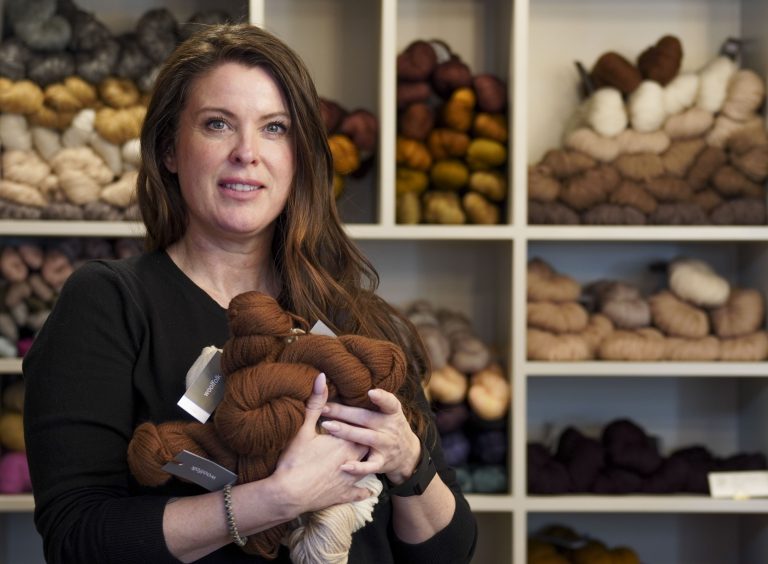
The Joy of Knitting – And Why It’s Time to Learn a Craft
Michèle Bowland learned to knit under the patient guidance of her nana Maureen. As a six-year-old, petting yarn,




